Posts: 307
Threads: 2
Joined: Sep 2023
04-08-2024, 02:29 PM
(This post was last modified: 04-08-2024, 02:30 PM by Kale.)
I've been thinking again about Basal Eurasian and the associated deep structure therein. Typically I run giant qpgraphs on the matter consisting of 24 populations to cover every base and make sure no false positives get through that would be shot down immediately with the introduction of one of these key populations.
Show Content
Spoiler
Mbuti
ZlatyKun
Ust_Ishim
BachoKiro_IUP
China_UP (Tianyuan & AR33k)
Andaman
Jomon
'East-Asian' (DevilsCave, etc.)
Native Americans
Kostenki14
Sunghir
Muierii
"Gravettian" (Vestonice, KremsWachtberg, Ostuni, Paglicci)
BachoKiro_BK1653
Goyet/Fournol
Yana
MA1
Georgia_UP (Kotias_UP, Dzudzuana)
Taforalt
Pinarbasi
CHG
Iran_N
WHG - Italian Epigravettian (AreneCandide, GrottaContinenza, etc.)
EHG
However, changes can be made to the deep structure without effecting certain other areas of the graph, that is to say some areas are self-contained or 'modular' as I've been referring to it in my own head as.
For example, whatever you do in the deeper areas of the graph will not change the very straightforward solution that Native Americans are East-Asian + ANE.
Having Native Americans in the full graph just helps constrain and focus the ANE component insofar as it contributes to other populations.
I've repeatedly asked for others to sketch graphs that illustrate their ideas, but have gotten 0 returns on that.
If we are discussing at the time depth of Basal Eurasian and initial peopling of Eurasia though, many of these 'modules' are largely irrelevant, and the list of 24 populations can be trimmed significantly, hopefully encouraging people to give their hand at working on this matter. These should be the basic 10 populations. Just make sure the MA1-Iran_N relationship is addressed by MA1 into Iran, not the other way around.
Mbuti
Iran_N
ZlatyKun
Ust_Ishim
BachoKiro_IUP
China_UP (Tianyuan & AR33k)
Andaman
Goyet/Fournol
Yana
MA1
Desdonas and Megalophias like this post
Posts: 73
Threads: 1
Joined: Oct 2023
(04-08-2024, 02:29 PM)Kale Wrote: I've been thinking again about Basal Eurasian and the associated deep structure therein. Typically I run giant qpgraphs on the matter consisting of 24 populations to cover every base and make sure no false positives get through that would be shot down immediately with the introduction of one of these key populations.
Show Content
Spoiler
Mbuti
ZlatyKun
Ust_Ishim
BachoKiro_IUP
China_UP (Tianyuan & AR33k)
Andaman
Jomon
'East-Asian' (DevilsCave, etc.)
Native Americans
Kostenki14
Sunghir
Muierii
"Gravettian" (Vestonice, KremsWachtberg, Ostuni, Paglicci)
BachoKiro_BK1653
Goyet/Fournol
Yana
MA1
Georgia_UP (Kotias_UP, Dzudzuana)
Taforalt
Pinarbasi
CHG
Iran_N
WHG - Italian Epigravettian (AreneCandide, GrottaContinenza, etc.)
EHG
However, changes can be made to the deep structure without effecting certain other areas of the graph, that is to say some areas are self-contained or 'modular' as I've been referring to it in my own head as.
For example, whatever you do in the deeper areas of the graph will not change the very straightforward solution that Native Americans are East-Asian + ANE.
Having Native Americans in the full graph just helps constrain and focus the ANE component insofar as it contributes to other populations.
I've repeatedly asked for others to sketch graphs that illustrate their ideas, but have gotten 0 returns on that.
If we are discussing at the time depth of Basal Eurasian and initial peopling of Eurasia though, many of these 'modules' are largely irrelevant, and the list of 24 populations can be trimmed significantly, hopefully encouraging people to give their hand at working on this matter. These should be the basic 10 populations. Just make sure the MA1-Iran_N relationship is addressed by MA1 into Iran, not the other way around.
Mbuti
Iran_N
ZlatyKun
Ust_Ishim
BachoKiro_IUP
China_UP (Tianyuan & AR33k)
Andaman
Goyet/Fournol
Yana
MA1
Do you think Anatolia/Caucasus and Kostenki/Gravettian are fully settled or why aren't they in the basic pops?
I've posted a graphlet like this before, what format do you want? This is mostly with your populations except with Kostenki instead of Goyet, surely Goyet should be the same thing with just some extra BachoKiro.
Show Content
Spoiler
Code: vertex R 0
vertex Mbuti.DG 0
vertex Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_bl 0
vertex Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_r 0
vertex Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_r_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_rh 0
vertex Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj 0
vertex Russia_Ust_Ishim_HG.DG 0
vertex admix 0
vertex admix_Russia_Kostenki14 0
vertex Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b 0
vertex Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrjwzv 0
vertex Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrjw 0
vertex admixb 0
vertex China_UP 0
vertex Russia_Kostenki14 0
vertex admixb_Russia_MA1_HG.SG 0
vertex Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrjwz 0
vertex Onge_Jarawa 0
vertex admixg 0
vertex Czechia_Bohemia_UP_HG 0
vertex Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_re 0
vertex Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_rh 0
vertex admixc 0
vertex Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_u 0
vertex Bulgaria_BachoKiro_LatePleistocene 0
vertex admixu 0
vertex Turkey_Epipaleolithic 0
vertex Russia_Yana_UP.SG 0
vertex admixb_Russia_MA1_HG.SGo 0
vertex Russia_MA1_HG.SG 0
vertex Iran_N 0
label Mbuti.DG Mbuti.DG
label Russia_Ust_Ishim_HG.DG Russia_Ust_Ishim_HG.DG
label China_UP China_UP
label Russia_Kostenki14 Russia_Kostenki14
label Onge_Jarawa Onge_Jarawa
label Czechia_Bohemia_UP_HG Czechia_Bohemia_UP_HG
label Bulgaria_BachoKiro_LatePleistocene Bulgaria_BachoKiro_LatePleistocene
label Turkey_Epipaleolithic Turkey_Epipaleolithic
label Russia_Yana_UP.SG Russia_Yana_UP.SG
label Russia_MA1_HG.SG Russia_MA1_HG.SG
label Iran_N Iran_N
edge R_Mbuti.DG R Mbuti.DG 0.028
edge R_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_bl R Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_bl 0.028
edge Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_bl_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_r Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_bl Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_r 0.002
edge Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_bl_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_r_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_rh Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_bl Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_r_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_rh 0.018
edge Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_Russia_Ust_Ishim_HG.DG Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj Russia_Ust_Ishim_HG.DG 0.003
edge admix_admix_Russia_Kostenki14 admix admix_Russia_Kostenki14 0.007
edge Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj 0.002
edge Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrjwzv Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrjwzv 0.004
edge Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrjw_China_UP Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrjw China_UP 0.016
edge admix_Russia_Kostenki14_Russia_Kostenki14 admix_Russia_Kostenki14 Russia_Kostenki14 0.139
edge admixb_admixb_Russia_MA1_HG.SG admixb admixb_Russia_MA1_HG.SG 0.003
edge Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrjwz_Onge_Jarawa Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrjwz Onge_Jarawa 0.012
edge Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_r_Czechia_Bohemia_UP_HG Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_r Czechia_Bohemia_UP_HG 0.15
edge Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_r_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_re Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_r Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_re 0.003
edge Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_r_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_rh_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_rh Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_r_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_rh Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_rh 0.057
edge Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_re_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_re Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b 0.001
edge Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_re_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_u Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_re Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_u 0.001
edge Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_u_Bulgaria_BachoKiro_LatePleistocene Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_u Bulgaria_BachoKiro_LatePleistocene 0.04
edge admixu_Turkey_Epipaleolithic admixu Turkey_Epipaleolithic 0.142
edge Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrjwzv_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrjw Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrjwzv Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrjw 0.004
edge Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrjwzv_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrjwz Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrjwzv Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrjwz 0.016
edge admixb_Russia_MA1_HG.SG_Russia_Yana_UP.SG admixb_Russia_MA1_HG.SG Russia_Yana_UP.SG 0.012
edge admixb_Russia_MA1_HG.SG_admixb_Russia_MA1_HG.SGo admixb_Russia_MA1_HG.SG admixb_Russia_MA1_HG.SGo 0.074
edge admixb_Russia_MA1_HG.SGo_Russia_MA1_HG.SG admixb_Russia_MA1_HG.SGo Russia_MA1_HG.SG 0.07
edge admixg_Iran_N admixg Iran_N 0.009
admix admix Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_rh 0.737 0.263
admix admixb Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrjw admix_Russia_Kostenki14 0.308 0.692
admix admixc Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_r_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_rh admixb_Russia_MA1_HG.SGo 0.961 0.039
admix admixg Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrjwz admixc 0.098 0.902
admix admixu Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_u Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_rh 0.558 0.442
Code: from to type weight low high
R Mbuti.DG edge 0.02831298009149051 NA NA
R Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_bl edge 0.028312980091406523 NA NA
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_bl Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_r edge 0.002178205754330613 NA NA
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_bl Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_r_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_rh edge 0.018171102099682312 NA NA
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj Russia_Ust_Ishim_HG.DG edge 0.0033391766174815474 NA NA
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj admix admix 0.7366939548382232 0.7366921880881088 0.7367899812815214
admix admix_Russia_Kostenki14 edge 0.006691327888981334 NA NA
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj edge 0.0015555911626918774 NA NA
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrjwzv edge 0.004300772856192601 NA NA
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrjw admixb admix 0.3078824721512031 0.30786262710148105 0.3078848756497642
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrjw China_UP edge 0.01597609955886178 NA NA
admix_Russia_Kostenki14 Russia_Kostenki14 edge 0.13941394313461986 NA NA
admix_Russia_Kostenki14 admixb admix 0.6921175278487969 0.6921151243502358 0.692137372898519
admixb admixb_Russia_MA1_HG.SG edge 0.0025666331011894783 NA NA
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrjwz Onge_Jarawa edge 0.012031671533008171 NA NA
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrjwz admixg admix 0.0982980915412315 0.0982677178354282 0.09834098267962796
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_r Czechia_Bohemia_UP_HG edge 0.1499376143308638 NA NA
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_r Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_re edge 0.002941420827987588 NA NA
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_r_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_rh Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_rh edge 0.05729094339313498 NA NA
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_r_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_rh admixc admix 0.9612830182541557 0.9612464330218091 0.9612830182541557
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_re Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b edge 8.614207354462657e-4 NA NA
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_re Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_u edge 7.041978569828527e-4 NA NA
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_u Bulgaria_BachoKiro_LatePleistocene edge 0.03954972735478601 NA NA
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_u admixu admix 0.5580143968909934 0.5580030014456366 0.5581657954085391
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_rh admix admix 0.2633060451617768 0.2632100187184786 0.26330781191189123
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_rh admixu admix 0.44198560310900664 0.44183420459146094 0.44199699855436336
admixu Turkey_Epipaleolithic edge 0.1424721245341429 NA NA
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrjwzv Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrjw edge 0.0036431982002490225 NA NA
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrjwzv Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrjwz edge 0.016149838221183786 NA NA
admixc admixg admix 0.9017019084587685 0.901659017320372 0.9017322821645718
admixb_Russia_MA1_HG.SG Russia_Yana_UP.SG edge 0.012016004978572564 NA NA
admixb_Russia_MA1_HG.SG admixb_Russia_MA1_HG.SGo edge 0.07447780446724689 NA NA
admixb_Russia_MA1_HG.SGo Russia_MA1_HG.SG edge 0.06954694022329186 NA NA
admixb_Russia_MA1_HG.SGo admixc admix 0.03871698174584426 0.03871698174584426 0.03875356697819088
admixg Iran_N edge 0.0093742779413086 NA NA
Code: digraph G {
label = "";
labelloc=t;
labeljust=l;
size = "7.5,10";
R [shape = point];
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_bl [shape = point];
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj [shape = point];
admix [shape = point];
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b [shape = point];
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrjw [shape = point];
admix_Russia_Kostenki14 [shape = point];
admixb [shape = point];
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrjwz [shape = point];
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_r [shape = point];
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_r_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_rh [shape = point];
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_re [shape = point];
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_u [shape = point];
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_rh [shape = point];
admixu [shape = point];
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrjwzv [shape = point];
admixc [shape = point];
admixb_Russia_MA1_HG.SG [shape = point];
admixb_Russia_MA1_HG.SGo [shape = point];
admixg [shape = point];
R -> MbutiDG [ label = "28" ];
R -> Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_bl [ label = "28" ];
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_bl -> Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_r [ label = "2" ];
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_bl -> Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_r_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_rh [ label = "18" ];
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj -> Russia_Ust_Ishim_HGDG [ label = "3" ];
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj -> admix [ style=dotted, label = "74%" ];
admix -> admix_Russia_Kostenki14 [ label = "7" ];
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b -> Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj [ label = "2" ];
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b -> Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrjwzv [ label = "4" ];
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrjw -> admixb [ style=dotted, label = "31%" ];
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrjw -> China_UP [ label = "16" ];
admix_Russia_Kostenki14 -> Russia_Kostenki14 [ label = "139" ];
admix_Russia_Kostenki14 -> admixb [ style=dotted, label = "69%" ];
admixb -> admixb_Russia_MA1_HGSG [ label = "3" ];
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrjwz -> Onge_Jarawa [ label = "12" ];
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrjwz -> admixg [ style=dotted, label = "10%" ];
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_r -> Czechia_Bohemia_UP_HG [ label = "150" ];
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_r -> Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_re [ label = "3" ];
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_r_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_rh -> Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_rh [ label = "57" ];
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_r_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_rh -> admixc [ style=dotted, label = "96%" ];
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_re -> Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b [ label = "1" ];
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_re -> Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_u [ label = "1" ];
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_u -> Bulgaria_BachoKiro_LatePleistocene [ label = "40" ];
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_u -> admixu [ style=dotted, label = "56%" ];
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_rh -> admix [ style=dotted, label = "26%" ];
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrj_b_Turkey_Epipaleolithic_rh -> admixu [ style=dotted, label = "44%" ];
admixu -> Turkey_Epipaleolithic [ label = "142" ];
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrjwzv -> Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrjw [ label = "4" ];
Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrjwzv -> Rr_Rrlr_Rr_Rrlr_Rr_Rrlr_Rrlrjwz [ label = "16" ];
admixc -> admixg [ style=dotted, label = "90%" ];
admixb_Russia_MA1_HGSG -> Russia_Yana_UPSG [ label = "12" ];
admixb_Russia_MA1_HGSG -> admixb_Russia_MA1_HGSGo [ label = "74" ];
admixb_Russia_MA1_HGSGo -> Russia_MA1_HGSG [ label = "70" ];
admixb_Russia_MA1_HGSGo -> admixc [ style=dotted, label = "4%" ];
admixg -> Iran_N [ label = "9" ];
}
The Iran/ANE relationship is a hard part because I think it might not be a simple one-way admixture, with more recent haplogroups X, R1a and R1b (mtDNA) shared between them.
Also, there's a plot_graph_map function in admixtools2, have you tried that? Maybe we could see big things with it, like the Persian plateau hub 
Megalophias likes this post
Posts: 307
Threads: 2
Joined: Sep 2023
04-08-2024, 05:31 PM
(This post was last modified: 04-08-2024, 06:04 PM by Kale.)
Pinarbasi/Caucasus_UP/Kostenki/Sunghir/Muierii/Gravettian, fall into a number of those 'modules'. You can do a number of things with them, but what will not change is that they all function well as something intermediate between Goyet and Iran_N (minus ANE). I know Chad is in agreement about the idea of Iran_N representing a significant pole in the Near-East/Basal/West arena of things. Not sure if he's also on board with Goyet being the opposite end of that spectrum.
In regards to sketching out ideas, I was more talking more about people that don't/can't run qpgraph, so I meant sketch literally if need be.
Hmm didn't know about plot_graph_map, I'll have to see what that looks like.
EDIT: Don't know what's going wrong with plot_graph_map
Error in plot_graph_map(graph, leafcoords):
all(c("iid", "group", "lat", "lon") %in% names(leafcoords)) is not TRUE
Maybe I misunderstand how this works? Do I have to supply coordinates for the mixture events or just the populations in the leafcoords file?
Posts: 73
Threads: 1
Joined: Oct 2023
(04-08-2024, 05:31 PM)Kale Wrote: Pinarbasi/Caucasus_UP/Kostenki/Sunghir/Muierii/Gravettian, fall into a number of those 'modules'. You can do a number of things with them, but what will not change is that they all function well as something intermediate between Goyet and Iran_N (minus ANE). I know Chad is in agreement about the idea of Iran_N representing a significant pole in the Near-East/Basal/West arena of things. Not sure if he's also on board with Goyet being the opposite end of that spectrum.
In regards to sketching out ideas, I was more talking more about people that don't/can't run qpgraph, so I meant sketch literally if need be.
Hmm didn't know about plot_graph_map, I'll have to see what that looks like.
EDIT: Don't know what's going wrong with plot_graph_map
Error in plot_graph_map(graph, leafcoords):
all(c("iid", "group", "lat", "lon") %in% names(leafcoords)) is not TRUE
Maybe I misunderstand how this works? Do I have to supply coordinates for the mixture events or just the populations in the leafcoords file? There's an example_anno object that shows what the format should be, I think it's best if you make one from the real anno file for just your populations with that four columns. But maybe not worth it as the result isn't really spectacular.

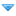
(04-08-2024, 12:28 AM)Desdonas Wrote: The significant bottleneck (over 10k) of C, F and D1 relative to E and D2 does indicate that the OoA population has a relatively isolated history (likely geographically). Of course, it is impossible to have an Arabian Standstill over 30k and an OoA of 80kya, but I believe it is reasonable for them to list these alleles. Perhaps this is a part of the OoA bottleneck itself, and if the population size is not large, the selection may complete quickly (during 10k?). Or we need more genome from East Africa to check whether pre-C and pre-F exist.
Second, Vallini's "Tianyuan-K14" binary model may indeed appear more "traditional" in 2024. But in terms of climate, the Iranian Plateau is indeed feasible as a hub for Eurasians, and the allele selection may also occur during the migration from East Africa to Iran (65-55kya).
Finally, Eurasians may include multiple components, including Crown Eurasians, ZK/para ZK, and a deeper BE. The third may come from the Arabian Peninsula, while the first two may both have roots in Iran (47kya and 50kya?). Para-ZK may be represented by y-hg G (also IJ?), while deep BE may be represented by some mt-N's primary branches. They have different split depth but may both contribute to West Eurasian populations. We are being diplomatic but that study is almost useless. Why not try to fit a tree onto their map like I just did, or at least calculate the allele sharing for a few more populations. What were they even thinking? "Let's gather the greatest scientists, we are calculating allele sharing with Kostenki and Tianyuan" 
They don't show that Iran was feasible as a hub, it was barely habitable, it's just that other regions like Arabia were even less so and they just wanted to choose a hub somewhere there. They actually suggest a decreased suitability for Iran in the period for their standstill.
At least they show that North India had a much higher sustainable population, makes it the right place for an East Eurasian hub, while the small West/Basal population was waiting for their big Heinrich event and Campanian eruption. Iran and the Central Asian desert was more like the divider between them, just like how it is today too. Occam says this two-way model is better than East+West+Basal+ZK+etc.
![[Image: 41467_2024_46161_Fig4_HTML.png?as=webp]](https://media.springernature.com/lw685/springer-static/image/art%3A10.1038%2Fs41467-024-46161-7/MediaObjects/41467_2024_46161_Fig4_HTML.png?as=webp)
Megalophias likes this post
Posts: 100
Threads: 5
Joined: Oct 2023
(04-07-2024, 04:44 PM)TanTin Wrote: Next, I will give you the same results for Denisova3-Neanderthal , but will use a different arrangement to correspond the time - space location for the above groups.
The arrangement is made by using another F4 statistics, that I am not showing here.
![[Image: Neand-Denisova-all-3.png]](https://i.postimg.cc/VvsdL06r/Neand-Denisova-all-3.png)
Thank you.
Btw could you add those various east Eurasian populations that you did with on page 1?
Posts: 100
Threads: 5
Joined: Oct 2023
(04-07-2024, 03:23 PM)TanTin Wrote: (04-07-2024, 11:28 AM)Jerome Wrote: Could you add Kostenki,afontovaGora,Tutkaul and Tyumen_HG here, dosent this show WHG as having more Neanderthal than MA1?
And iran_n has more Neanderthal affinity(2.16) than ust ishim(1.87)?
Dosent this refute lazaridis conception of basal eurasian as needed to account for lower Neanderthal Ancestry?
Maybe deselection against Neanderthal alleles could explain that??
Why else is basal eurasian even needed now?
There's also the fact that iran and central Asia was full of Neanderthals(warsawi,Dodekatym-2) so the area being full of basal eurasians dosent make sense.
Any basal eurasian living there would end up with lots of Neanderthal Ancestry which defeats the purpose of basal itself.
I did the rerun.
Show Content
Spoiler
pop1 pop2 pop3 pop4 est se z p n
<chr> <chr> <chr> <chr> <dbl> <dbl> <dbl> <dbl> <dbl>
1 Primate_Chimp Altai_Neanderthal.DG Yoruba Primate_Chimp -0.0959 0.000530 -181. 0 592996
2 Primate_Chimp Altai_Neanderthal.DG Yoruba Altai_Neanderthal.DG 0.0798 0.000570 140. 0 592996
3 Primate_Chimp Altai_Neanderthal.DG Yoruba Russia_Ust_Ishim.DG 0.00168 0.000352 4.76 1.96e- 6 592969
4 Primate_Chimp Altai_Neanderthal.DG Yoruba Russia_MA1_HG.SG 0.00181 0.000358 5.05 4.41e- 7 425966
5 Primate_Chimp Altai_Neanderthal.DG Yoruba Luxembourg_Loschbour.DG 0.00156 0.000290 5.39 6.87e- 8 592749
6 Primate_Chimp Altai_Neanderthal.DG Yoruba Bulgaria_N 0.00146 0.000268 5.44 5.20e- 8 508318
7 Primate_Chimp Altai_Neanderthal.DG Yoruba Iran_GanjDareh_N 0.00117 0.000231 5.05 4.39e- 7 562236
8 Primate_Chimp Altai_Neanderthal.DG Yoruba Iran_HajjiFiruz_N 0.00139 0.000247 5.62 1.96e- 8 539981
9 Primate_Chimp Altai_Neanderthal.DG Yoruba Iranian 0.00133 0.000177 7.52 5.30e-14 592996
10 Primate_Chimp Altai_Neanderthal.DG Yoruba Mozabite 0.00125 0.000159 7.88 3.18e-15 592996
11 Primate_Chimp Altai_Neanderthal.DG Yoruba Georgia_Kotias.SG 0.00162 0.000298 5.43 5.70e- 8 591680
12 Primate_Chimp Altai_Neanderthal.DG Yoruba Georgia_Satsurblia.SG 0.00113 0.000365 3.09 1.98e- 3 352936
13 Primate_Chimp Altai_Neanderthal.DG Yoruba Ju_hoan_North 0.00116 0.000170 6.82 9.27e-12 592996
14 Primate_Chimp Altai_Neanderthal.DG Yoruba Egyptian 0.00111 0.000160 6.95 3.71e-12 592996
15 Primate_Chimp Altai_Neanderthal.DG Yoruba Israel_Natufian 0.000790 0.000522 1.51 1.30e- 1 115788
16 Primate_Chimp Altai_Neanderthal.DG Yoruba Israel_Natufian_contam 0.00138 0.000403 3.43 5.94e- 4 249108
17 Primate_Chimp Altai_Neanderthal.DG Yoruba Algerian 0.00106 0.000167 6.36 1.97e-10 592996
18 Primate_Chimp Altai_Neanderthal.DG Yoruba South_Africa_400BP.SG 0.000189 0.000159 1.19 2.36e- 1 592957
19 Primate_Chimp Altai_Neanderthal.DG Yoruba South_Africa_1900BP.SG 0.000863 0.000249 3.47 5.15e- 4 592913
20 Primate_Chimp Altai_Neanderthal.DG Yoruba Saharawi 0.000907 0.000164 5.52 3.46e- 8 592996
1 Primate_Chimp Altai_Neanderthal.DG Yoruba Yoruba 0 0 NaN NaN 592996
2 Primate_Chimp Altai_Neanderthal.DG Yoruba JPT.SG 0.00159 0.000223 7.13 1.04e-12 592995
3 Primate_Chimp Altai_Neanderthal.DG Yoruba Malawi_Hora_LSA_15500BP 0.000266 0.000571 0.466 6.41e- 1 105546
4 Primate_Chimp Altai_Neanderthal.DG Yoruba Malawi_Hora_LSA_8500BP 0.00130 0.000956 1.36 1.72e- 1 37456
5 Primate_Chimp Altai_Neanderthal.DG Yoruba Malawi_Fingira_LSA_6000BP 0.000686 0.000438 1.57 1.17e- 1 186016
6 Primate_Chimp Altai_Neanderthal.DG Yoruba Malawi_Fingira_LSA_2500BP 0.000403 0.000294 1.37 1.70e- 1 534105
7 Primate_Chimp Altai_Neanderthal.DG Yoruba Malawi_Chencherere_LSA_5200BP 0.000401 0.000520 0.771 4.40e- 1 126318
8 Primate_Chimp Altai_Neanderthal.DG Yoruba Russia_Kostenki14 0.00173 0.000349 4.95 7.32e- 7 584621
9 Primate_Chimp Altai_Neanderthal.DG Yoruba Russia_AfontovaGora3 0.00169 0.000517 3.27 1.09e- 3 163392
10 Primate_Chimp Altai_Neanderthal.DG Yoruba Russia_AfontovaGora2.SG 0.00111 0.000830 1.34 1.81e- 1 48428
11 Primate_Chimp Altai_Neanderthal.DG Yoruba Russia_Tyumen_HG 0.00209 0.000353 5.92 3.28e- 9 443261
![[Image: F4-Kostenki.png]](https://i.postimg.cc/R0SZFL6M/F4-Kostenki.png)
Are these scores based on z value or the weights?
Posts: 100
Threads: 5
Joined: Oct 2023
Does anyone know if deselection against Neanderthal alleles can lower the apparent allele sharing?
Posts: 71
Threads: 0
Joined: Nov 2023
04-09-2024, 03:19 AM
(This post was last modified: 04-09-2024, 03:26 AM by Desdonas.)
@ Kale @ kolompar
What do you think of India as a hub for early Eurasian population? I have seen that many haplogroup migration maps locate the origin of F in India, and India seems to have some unresolved F*. If this would be true, the main OoA bottleneck could be in there. IJ and G (west/basal) headed west first, then C, D1, F2, H and K stayed here and shared extra bottleneck for about 3,000 years. India is on a similar temperature zone of the Arabian peninsula, and the OoA allele selection may be possible.
However, if India is not the hub, then there may be some issues on the "Arabian Standstill". Do you agree that this selection/OoA bottleneck had already happened within Africa, and there was no real standstill after the exodus out of Africa (55-50kya)? If so, maybe all OoA related elements (pre-C, pre-F, pre-D1, etc.) were taken over by the E-population later.
Posts: 100
Threads: 5
Joined: Oct 2023
I think it was more likely the regions around Caspian sea and western central Asia.
From there you can easily head to Caucasus and Anatolia on one way and central Asia - Siberia on other,giving rise to early IUP industries like kulbalkian.
South Asia is less likely, because of its late IUP/EUP industries in centre.
They seem to be mostly present in south or in NW,where they are connected with central Asian industries.
Posts: 307
Threads: 2
Joined: Sep 2023
04-09-2024, 04:04 AM
(This post was last modified: 04-09-2024, 04:05 AM by Kale.)
(04-09-2024, 03:19 AM)Desdonas Wrote: @Kale @kolompar
What do you think of India as a hub for early Eurasian population? I have seen that many haplogroup migration maps locate the origin of F in India, and India seems to have some unresolved F*. If this would be true, the main OoA bottleneck could be in there. IJ and G (west/basal) headed west first, then C, D1, F2, H and K stayed here and shared extra bottleneck for about 3,000 years. India is on a similar temperature zone of the Arabian peninsula, and the OoA allele selection may be possible.
However, if India is not the hub, then there may be some issues on the "Arabian Standstill". Do you agree that this selection/OoA bottleneck had already happened within Africa, and there was no real standstill after the exodus out of Africa (55-50kya)? If so, maybe all OoA related elements (pre-C, pre-F, pre-D1, etc.) were taken over by the E-population later.
Modern distribution seems to mean less and less the more ancient samples we get. Imagine if BachoKiro's pre-F2'4, Kostenki's C1b, Oase's K2a-M2308 and mt-pre-N, Goyet & Ostuni's Mt-M, etc. all survived? I think India and Southeast Asia somehow just preserved more early diversity, not that they are the source of it.
In regards to rapid OOA vs Arabian standstill, I haven't looked into the idea enough to really form a perception.
Merriku and Megalophias like this post
Posts: 14
Threads: 0
Joined: Oct 2023
Gender: Male
Ethnicity: Chhattisgarhi
Nationality: Indian
Y-DNA (P): L-Y46452 > L-Y266345 (L1a2)
mtDNA (M): M2a3c*
(04-09-2024, 03:19 AM)Desdonas Wrote: @Kale @kolompar
What do you think of India as a hub for early Eurasian population? I have seen that many haplogroup migration maps locate the origin of F in India, and India seems to have some unresolved F*. If this would be true, the main OoA bottleneck could be in there. IJ and G (west/basal) headed west first, then C, D1, F2, H and K stayed here and shared extra bottleneck for about 3,000 years. India is on a similar temperature zone of the Arabian peninsula, and the OoA allele selection may be possible.
However, if India is not the hub, then there may be some issues on the "Arabian Standstill". Do you agree that this selection/OoA bottleneck had already happened within Africa, and there was no real standstill after the exodus out of Africa (55-50kya)? If so, maybe all OoA related elements (pre-C, pre-F, pre-D1, etc.) were taken over by the E-population later.
I am skeptical of those Indian Fs. Considering the amount of Indian private samples I think F should have turned up by now.
I think those Indian F*s are from old (STR based?) studies. If I am not mistaken, when these studies were published haplogroup H3 wasn't known & many of those F* may actually be H3.
I don't know about FTDNA but both theytree and yfull don't have any Indian (F ×GHIJK) samples.
The only clue we have is 2 F2 samples from the user Tipirneni's 23&me matches , but then again it could just be the case of older nomenclature.
Posts: 102
Threads: 2
Joined: Sep 2023
Gender: Male
Ethnicity: Finn
Nationality: Finnish
Y-DNA (P): I1-L22
Y-DNA (M): J1c7a
That Ju-Hoan stat is very interesting
I have three propositions
Either the South Africans have genuine Neanderthal ancestry, they have some deeper Denisovan related ancestry or the Human ancestry in Neanderthals is closest to South Africans in particular.
Today this wouldn't make sense, but there's always a chance South Africans used to reside closer to Neanderthals than the ancestors of all other Humans did.
The low stat for West Africans can be explained with archaic ancestry, but South Africans are higher than many Eurasians are.
Posts: 363
Threads: 8
Joined: Oct 2023
(04-09-2024, 01:40 AM)Jerome Wrote: (04-07-2024, 03:23 PM)TanTin Wrote: (04-07-2024, 11:28 AM)Jerome Wrote: Could you add Kostenki,afontovaGora,Tutkaul and Tyumen_HG here, dosent this show WHG as having more Neanderthal than MA1?
And iran_n has more Neanderthal affinity(2.16) than ust ishim(1.87)?
Dosent this refute lazaridis conception of basal eurasian as needed to account for lower Neanderthal Ancestry?
Maybe deselection against Neanderthal alleles could explain that??
Why else is basal eurasian even needed now?
There's also the fact that iran and central Asia was full of Neanderthals(warsawi,Dodekatym-2) so the area being full of basal eurasians dosent make sense.
Any basal eurasian living there would end up with lots of Neanderthal Ancestry which defeats the purpose of basal itself.
I did the rerun.
Show Content
Spoiler
pop1 pop2 pop3 pop4 est se z p n
<chr> <chr> <chr> <chr> <dbl> <dbl> <dbl> <dbl> <dbl>
1 Primate_Chimp Altai_Neanderthal.DG Yoruba Primate_Chimp -0.0959 0.000530 -181. 0 592996
2 Primate_Chimp Altai_Neanderthal.DG Yoruba Altai_Neanderthal.DG 0.0798 0.000570 140. 0 592996
3 Primate_Chimp Altai_Neanderthal.DG Yoruba Russia_Ust_Ishim.DG 0.00168 0.000352 4.76 1.96e- 6 592969
4 Primate_Chimp Altai_Neanderthal.DG Yoruba Russia_MA1_HG.SG 0.00181 0.000358 5.05 4.41e- 7 425966
5 Primate_Chimp Altai_Neanderthal.DG Yoruba Luxembourg_Loschbour.DG 0.00156 0.000290 5.39 6.87e- 8 592749
6 Primate_Chimp Altai_Neanderthal.DG Yoruba Bulgaria_N 0.00146 0.000268 5.44 5.20e- 8 508318
7 Primate_Chimp Altai_Neanderthal.DG Yoruba Iran_GanjDareh_N 0.00117 0.000231 5.05 4.39e- 7 562236
8 Primate_Chimp Altai_Neanderthal.DG Yoruba Iran_HajjiFiruz_N 0.00139 0.000247 5.62 1.96e- 8 539981
9 Primate_Chimp Altai_Neanderthal.DG Yoruba Iranian 0.00133 0.000177 7.52 5.30e-14 592996
10 Primate_Chimp Altai_Neanderthal.DG Yoruba Mozabite 0.00125 0.000159 7.88 3.18e-15 592996
11 Primate_Chimp Altai_Neanderthal.DG Yoruba Georgia_Kotias.SG 0.00162 0.000298 5.43 5.70e- 8 591680
12 Primate_Chimp Altai_Neanderthal.DG Yoruba Georgia_Satsurblia.SG 0.00113 0.000365 3.09 1.98e- 3 352936
13 Primate_Chimp Altai_Neanderthal.DG Yoruba Ju_hoan_North 0.00116 0.000170 6.82 9.27e-12 592996
14 Primate_Chimp Altai_Neanderthal.DG Yoruba Egyptian 0.00111 0.000160 6.95 3.71e-12 592996
15 Primate_Chimp Altai_Neanderthal.DG Yoruba Israel_Natufian 0.000790 0.000522 1.51 1.30e- 1 115788
16 Primate_Chimp Altai_Neanderthal.DG Yoruba Israel_Natufian_contam 0.00138 0.000403 3.43 5.94e- 4 249108
17 Primate_Chimp Altai_Neanderthal.DG Yoruba Algerian 0.00106 0.000167 6.36 1.97e-10 592996
18 Primate_Chimp Altai_Neanderthal.DG Yoruba South_Africa_400BP.SG 0.000189 0.000159 1.19 2.36e- 1 592957
19 Primate_Chimp Altai_Neanderthal.DG Yoruba South_Africa_1900BP.SG 0.000863 0.000249 3.47 5.15e- 4 592913
20 Primate_Chimp Altai_Neanderthal.DG Yoruba Saharawi 0.000907 0.000164 5.52 3.46e- 8 592996
1 Primate_Chimp Altai_Neanderthal.DG Yoruba Yoruba 0 0 NaN NaN 592996
2 Primate_Chimp Altai_Neanderthal.DG Yoruba JPT.SG 0.00159 0.000223 7.13 1.04e-12 592995
3 Primate_Chimp Altai_Neanderthal.DG Yoruba Malawi_Hora_LSA_15500BP 0.000266 0.000571 0.466 6.41e- 1 105546
4 Primate_Chimp Altai_Neanderthal.DG Yoruba Malawi_Hora_LSA_8500BP 0.00130 0.000956 1.36 1.72e- 1 37456
5 Primate_Chimp Altai_Neanderthal.DG Yoruba Malawi_Fingira_LSA_6000BP 0.000686 0.000438 1.57 1.17e- 1 186016
6 Primate_Chimp Altai_Neanderthal.DG Yoruba Malawi_Fingira_LSA_2500BP 0.000403 0.000294 1.37 1.70e- 1 534105
7 Primate_Chimp Altai_Neanderthal.DG Yoruba Malawi_Chencherere_LSA_5200BP 0.000401 0.000520 0.771 4.40e- 1 126318
8 Primate_Chimp Altai_Neanderthal.DG Yoruba Russia_Kostenki14 0.00173 0.000349 4.95 7.32e- 7 584621
9 Primate_Chimp Altai_Neanderthal.DG Yoruba Russia_AfontovaGora3 0.00169 0.000517 3.27 1.09e- 3 163392
10 Primate_Chimp Altai_Neanderthal.DG Yoruba Russia_AfontovaGora2.SG 0.00111 0.000830 1.34 1.81e- 1 48428
11 Primate_Chimp Altai_Neanderthal.DG Yoruba Russia_Tyumen_HG 0.00209 0.000353 5.92 3.28e- 9 443261
![[Image: F4-Kostenki.png]](https://i.postimg.cc/R0SZFL6M/F4-Kostenki.png)
Are these scores based on z value or the weights?
These are the F4. The first column - for estimation.
The vertical intervals are for the error.
Z is not shown on the picture, but it is in the table in the 3-rd column.
Z = est/se . se is the standard error.
Posts: 363
Threads: 8
Joined: Oct 2023
(04-09-2024, 11:05 AM)Norfern-Ostrobothnian Wrote: That Ju-Hoan stat is very interesting
I have three propositions
Either the South Africans have genuine Neanderthal ancestry, they have some deeper Denisovan related ancestry or the Human ancestry in Neanderthals is closest to South Africans in particular.
Today this wouldn't make sense, but there's always a chance South Africans used to reside closer to Neanderthals than the ancestors of all other Humans did.
The low stat for West Africans can be explained with archaic ancestry, but South Africans are higher than many Eurasians are.
I did additional tests yesterday to find where this Neanderthal component came for South Africans.
It is not genuine Neanderthal.
They admixed with some North African archaic group which had already the Neanderthal components.
And that's why they score high .
Before to expand to Europe and Asia, the Neanderthals were obviously in North Africa.
Posts: 102
Threads: 2
Joined: Sep 2023
Gender: Male
Ethnicity: Finn
Nationality: Finnish
Y-DNA (P): I1-L22
Y-DNA (M): J1c7a
The level is so high that it'd require the likely East African Nilotic or Cushitic ancestry to be a very large portion of their ancestry. Either this is artefacting, which it might be, or this is genuine ancestry from Neanderthal related populations.
|






![[Image: 41467_2024_46161_Fig4_HTML.png?as=webp]](https://media.springernature.com/lw685/springer-static/image/art%3A10.1038%2Fs41467-024-46161-7/MediaObjects/41467_2024_46161_Fig4_HTML.png?as=webp)
![[Image: Neand-Denisova-all-3.png]](https://i.postimg.cc/VvsdL06r/Neand-Denisova-all-3.png)
![[Image: F4-Kostenki.png]](https://i.postimg.cc/R0SZFL6M/F4-Kostenki.png)